UPDATE: I built a tool to solve some of the challenges mentioned in the post. It’s now live at https://www.reviewnb.com/
————————————————————————————————————————————–
A lot of people, including me, love Jupyter Notebooks. It’s a fantastic tool for data science. Today I’m not going to talk about it’s amazing capabilities but rather how it fails at two important things: Version Control and Reproducibility. I will also outline the current state-of-the-art to solve these problems. It’s a useful read if you are a Jupyter user. Let’s jump right in.
Version Control
Jupyter Notebook renders nicely in the browser and shows code, markdown and output in a single document. Behind the scenes, notebook file is a large JSON blob that looks like this. The format makes it hard to get notebooks to work with version control system like Git. Let’s look at two important version control workflows: Diff & Merge.
- Diffs are hard to read
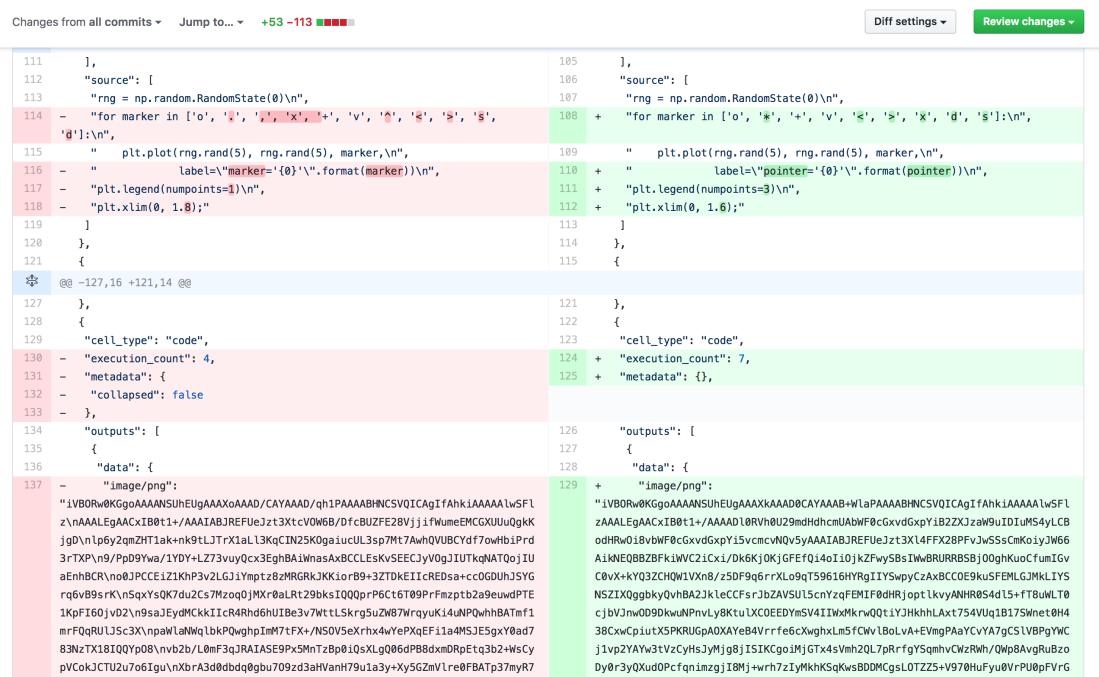
Notebook JSON includes everything: raw markdown, code snippets, metadata, and even output images represented as binary strings. As you can see in the above picture, diffs are pretty hard to read (GitHub PR link). Images can’t be compared, code is inside JSON arrays, lot of irrelevant metadata changes and so on. It’s impossible to review PR with this kind of diff.
Tools like nbdime are created to see diff in more human friendly way. But nbdime is only useful locally and don’t work with version control systems like GitHub/GitLab that people actually use. We need a way to look at proper diff in GitHub Pull Request and be able to comment etc.
- Merge is even harder
Let’s say you somehow manage to do without diff. Now your teammate has pushed changes to a notebook and you wish to pull those in and merge with your local changes. Good luck resolving the merge conflict in any text based editor. You have to take care of JSON format integrity, image binary strings, numerous metadata changes and so on.
Users typically fall back on –theirs/–ours git strategy since the effort to actually merge is too high. You can also setup nbmerge as git driver to manually resolve merge conflicts inside the Jupyter UI. This is definitely better than mucking around the JSON in a text editor.
Reproducibility
Notebook code (any code for that matter) requires certain environment to work correctly. This includes other packages, environment variables, data files etc. Jupyter, by design, doesn’t capture the environment information anywhere. Given just Notebook files, it’s not always easy or possible to reproduce the result.
Check out the BinderHub project for creating reproducible notebook environments. You provide it a GitHub URL, it looks for requirements.txt or environment.yml file, builds a docker image & spins up a JupyterHub server with your notebooks in it. The project is still in beta but might be your best bet for reproducible notebook environments at the moment.
Conclusion
In all fairness, Jupyter was designed for individual use but given the simplicity & popularity it’s getting adopted by teams for sharing/collaboration workflow as well. Certain design choices, probably made at its inception, don’t bode well with requirements of team workflow.
That’s all. I’m also working on a tool that solves the version control problem and makes it easy to use notebooks with GitHub (and likely with GitLab later).
UPDATE: The tool is now live at https://www.reviewnb.com/
One thought on “Challenges with Jupyter Notebook”